


Lineage Frequency Dynamics and Growth-Advantage Estimation from Genomic Surveillance Counts
An R package for modeling pathogen lineage frequencies, estimating growth advantages, and forecasting variant replacement dynamics from genomic surveillance counts.

Three lines of code transform raw surveillance counts into publication-ready model fits, growth advantage estimates, and probabilistic forecasts — with built-in backtesting for honest accuracy evaluation.
| Without lineagefreq | With lineagefreq |
|---|---|
| Raw point estimates, no model | MLR / hierarchical MLR / Piantham engines |
| No uncertainty quantification | 95% prediction intervals (parameter + sampling) |
| No forecasting | Probabilistic 2–6 week frequency forecasts |
| No evaluation framework | Rolling-origin backtest + MAE/WIS/coverage |
| Ad hoc scripts per analysis | Reproducible lfq_data → fit_model →
forecast pipeline |
| Not on CRAN | CRAN-distributable, tested on 4 platforms |
# install.packages("pak")
pak::pak("CuiweiG/lineagefreq")
# Or with devtools:
# devtools::install_github("CuiweiG/lineagefreq")library(lineagefreq)
library(ggplot2)
data(cdc_sarscov2_jn1)
x <- lfq_data(cdc_sarscov2_jn1,
lineage = lineage, date = date, count = count)
fit <- fit_model(x, engine = "mlr")
growth_advantage(fit, type = "relative_Rt", generation_time = 5)
fc <- forecast(fit, horizon = 28)
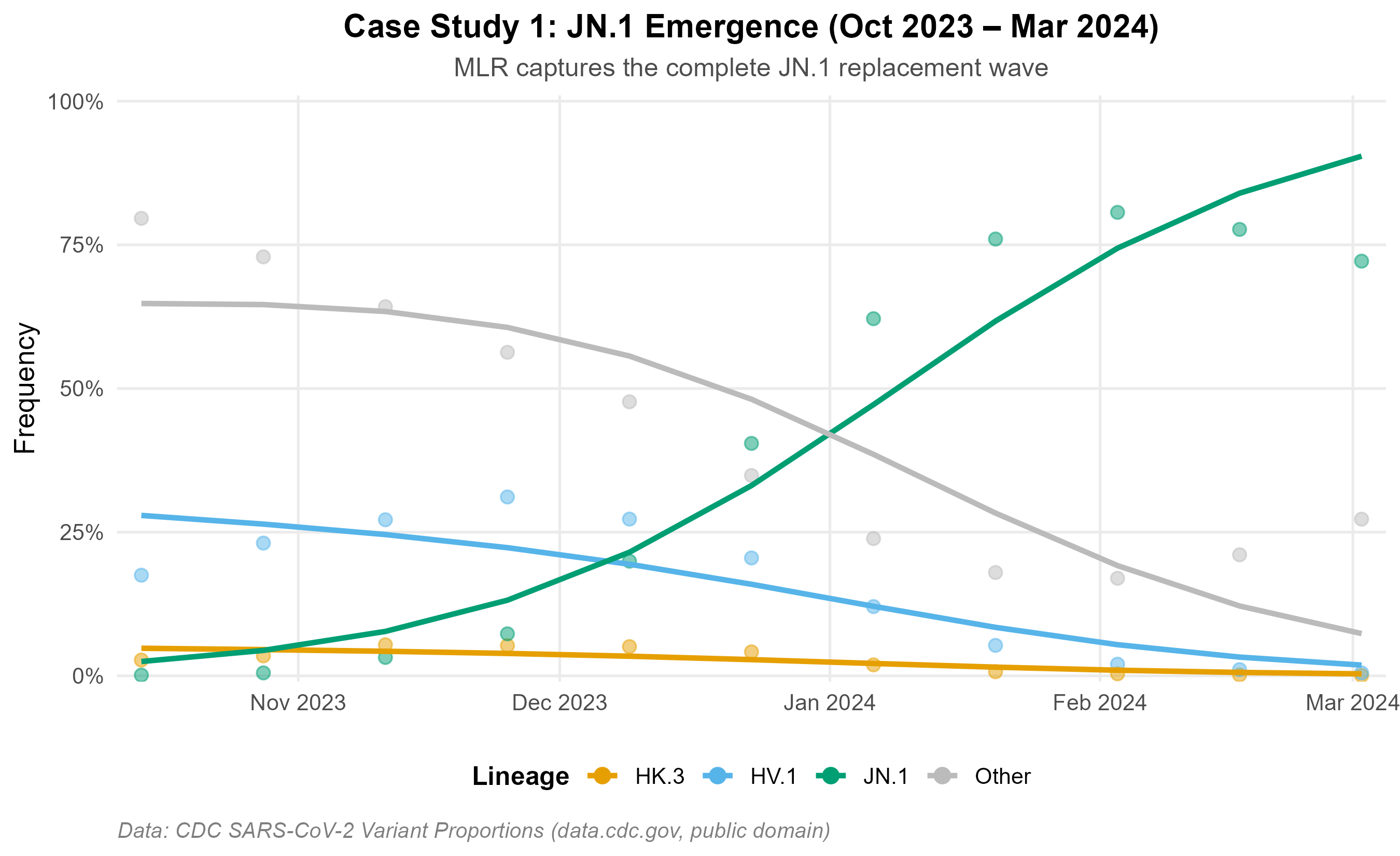
autoplot(fc)Figures below use real U.S. CDC surveillance data (data.cdc.gov/jr58-6ysp, public domain). Two independent epidemic waves illustrate model behavior across distinct replacement settings.
Data accessed 2026-03-28. Lineages below 5% peak frequency collapsed
to “Other.” Reproducible scripts:
data-raw/prepare_cdc_data.R and
data-raw/prepare_ba2_data.R.
JN.1 emergence (Oct 2023 – Mar 2024): MLR recovers the observed replacement trajectory from <1% to >80%.
BA.1 → BA.2 period (Dec 2021 – Jun 2022): A well-characterized Omicron replacement wave with four sequential subvariant sweeps.

Relative Rt estimates are consistent with published values: BA.2 = 1.34× vs BA.1 (Lyngse et al. 2022, published 1.3–1.5×); KP.3 = 1.36× vs JN.1. Generation times: 3.2 days for Omicron BA.* subvariants (Du et al. 2022); 5.0 days for JN/KP lineages.

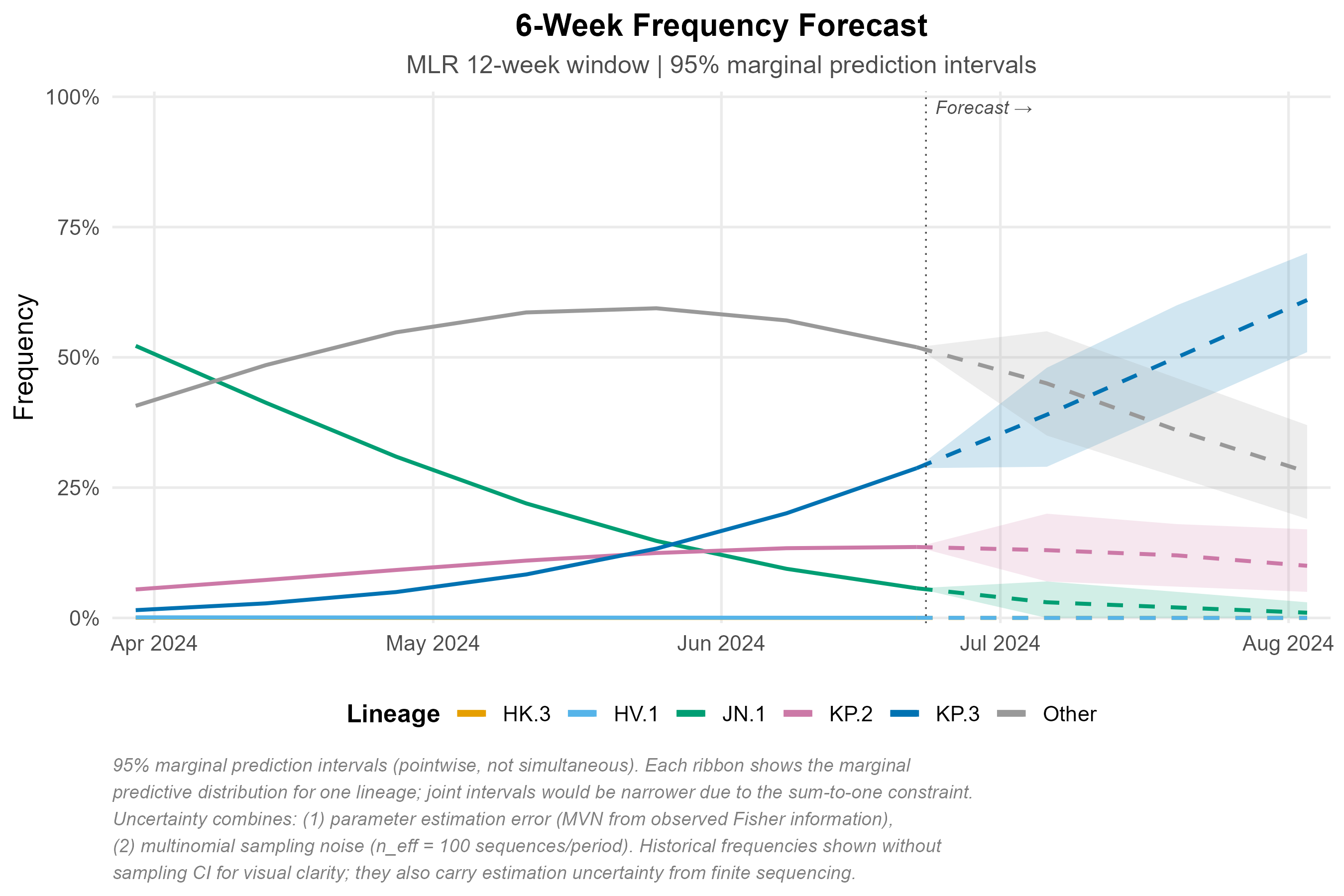
Six-week projection with 95% marginal prediction intervals (pointwise, not simultaneous). Uncertainty reflects parameter estimation error (MVN from Fisher information) and multinomial sampling noise (n_eff = 100 sequences/period). See figure caption for full methodological notes.
Rolling-origin out-of-sample evaluation on the BA.2 period: approximately 4% MAE at 2-week and 8% at 4-week horizon.

Model fitting - fit_model() with
engines "mlr", "hier_mlr",
"piantham", "fga", "garw"
(Bayesian engines require ‘CmdStan’)
Inference - Growth advantage in four scales: growth rate, relative Rt, selection coefficient, doubling time
Forecasting - Probabilistic frequency forecasts with parametric simulation and configurable sampling noise
Evaluation - Rolling-origin backtesting via
backtest() with standardized scoring (MAE, RMSE, coverage,
WIS) via score_forecasts()
Surveillance utilities -
summarize_emerging(): binomial GLM trend tests per lineage
- sequencing_power(): minimum sample size for detection -
collapse_lineages(), filter_sparse():
preprocessing
Visualization - autoplot() methods for
fits, forecasts, and backtest summaries - Publication-quality output
with colorblind-safe palettes
Interoperability - broom-compatible:
tidy(), glance(), augment() -
as_lfq_data() generic for extensible data import -
read_lineage_counts() for CSV input
Any pathogen with variant/lineage-resolved sequencing count data: SARS-CoV-2, influenza, RSV, mpox, and others.
citation("lineagefreq")A software paper and Zenodo DOI will be added upon publication.
MIT