{mnirs} contains standardised, reproducible methods for reading, processing, and analysing data from muscle near-infrared spectroscopy (mNIRS) devices. Intended for mNIRS researchers and practitioners in exercise physiology, sports science, and clinical rehabilitation with minimal coding experience required.
install.packages("mnirs")You can install the development version of
mnirs from GitHub with:
# install.packages("pak")
pak::pak("jemarnold/mnirs")<coming soon>
A very basic implementation of this package is hosted at https://jemarnold-mnirs-app.share.connect.posit.cloud/ and can currently be used for reading and pre-processing mNIRS data.
A more detailed vignette for common usage can be found on the package website: Reading and Cleaning Data with {mnirs}
mnirs is designed for mNIRS data, but it can be
used to read, clean, and process other time series datasets which
require many of the same processing steps. Enjoy!
read_mnirs() Read
data from filelibrary(ggplot2) ## for plotting
library(patchwork) ## for plotting
library(mnirs)
## {mnirs} includes sample files from a few mNIRS devices
example_mnirs()
#> [1] "artinis_intervals.xlsx" "moxy_intervals.csv"
#> [3] "moxy_ramp.xlsx" "portamon-oxcap.xlsx"
#> [5] "train.red_intervals.csv"
## rename channels in the format `renamed = "original_name"`
## where "original_name1" should match the file column name exactly
data_raw <- read_mnirs(
file_path = example_mnirs("moxy_ramp"), ## call an example data file
nirs_channels = c(
smo2_left = "SmO2 Live", ## identify and rename channels
smo2_right = "SmO2 Live(2)"
),
time_channel = c(time = "hh:mm:ss"), ## date-time format will be converted to numeric
event_channel = NULL, ## leave blank if unused
sample_rate = NULL, ## if blank, will be estimated from time_channel
add_timestamp = FALSE, ## omit a date-time timestamp column
zero_time = TRUE, ## recalculate time values from zero
keep_all = FALSE, ## return only the specified data channels
verbose = TRUE ## show warnings & messages
)
#> ! Estimated `sample_rate` = 2 Hz.
#> ℹ Define `sample_rate` explicitly to override.
#> Warning: ! Duplicate or irregular `time_channel` samples detected.
#> ℹ Investigate at `time` = 211.99 and 1184.
#> ℹ Re-sample with `mnirs::resample_mnirs()`.
## Note the above info message that sample_rate was estimated correctly at 2 Hz 👆
## ignore the warnings about irregular sampling for now, we will resample later
data_raw
#> # A tibble: 2,203 × 3
#> time smo2_left smo2_right
#> <dbl> <dbl> <dbl>
#> 1 0 54 68
#> 2 0.400 54 68
#> 3 0.960 54 68
#> 4 1.51 54 66
#> 5 2.06 54 66
#> 6 2.61 54 66
#> 7 3.16 54 66
#> 8 3.71 57 67
#> 9 4.26 57 67
#> 10 4.81 57 67
#> # ℹ 2,193 more rows
## note the `time_labels` plot argument to display time values as `h:mm:ss`
plot(
data_raw,
time_labels = TRUE,
n.breaks = 5,
na.omit = FALSE
)
mnirs data frames## view metadata
attr(data_raw, "nirs_channels")
#> [1] "smo2_left" "smo2_right"
attr(data_raw, "time_channel")
#> [1] "time"
attr(data_raw, "sample_rate")
#> [1] 2resample_mnirs():
Resample datadata_resampled <- resample_mnirs(
data_raw, ## blank channels will be retrieved from metadata
time_channel = time, ## channels can be left blank or specified explicitly
sample_rate = NULL, ## blank by default will be retrieved from metadata
resample_rate = 2, ## blank by default will resample to `sample_rate`
method = "linear" ## linear interpolation across resampled indices
)
#> ℹ Output is resampled at 2 Hz.
## note the altered "time" values from the original data frame 👇
data_resampled
#> # A tibble: 2,419 × 3
#> time smo2_left smo2_right
#> <dbl> <dbl> <dbl>
#> 1 0 54 68
#> 2 0.5 54 68
#> 3 1 54 67.9
#> 4 1.5 54 66.0
#> 5 2 54 66
#> 6 2.5 54 66
#> 7 3 54 66
#> 8 3.5 55.9 66.6
#> 9 4 57 67
#> 10 4.5 57 67
#> # ℹ 2,409 more rowsreplace_mnirs:
Replace local outliers, invalid values, and missing valuesdata_cleaned <- replace_mnirs(
data_resampled, ## blank channels will be retrieved from metadata
invalid_values = 0, ## known invalid values in the data
invalid_above = 90, ## remove data spikes above 90
outlier_cutoff = 3, ## recommended default value
width = 7, ## window to detect and replace outliers/missing values
method = "linear" ## linear interpolation over `NA`s
)
plot(data_cleaned, time_labels = TRUE)
filter_mnirs():
Digital filteringdata_filtered <- filter_mnirs(
data_cleaned, ## blank channels will be retrieved from metadata
method = "butterworth", ## Butterworth digital filter is a common choice
order = 2, ## filter order number
W = 0.02, ## filter fractional critical frequency `[0, 1]`
type = "low", ## specify a "low-pass" filter
na.rm = TRUE ## explicitly ignore NAs
)
## we will add the non-filtered data back to the plot to compare
plot(data_filtered, time_labels = TRUE) +
geom_line(
data = data_cleaned,
aes(y = smo2_left, colour = "smo2_left"), alpha = 0.4
) +
geom_line(
data = data_cleaned,
aes(y = smo2_right, colour = "smo2_right"), alpha = 0.4
)
shift_mnirs()
& rescale_mnirs(): Shift and rescale datadata_shifted <- shift_mnirs(
data_filtered, ## un-grouped nirs channels to shift separately
nirs_channels = list(smo2_left, smo2_right),
to = 0, ## NIRS values will be shifted to zero
span = 120, ## shift the *first* 120 sec of data to zero
position = "first"
)
plot(data_shifted, time_labels = TRUE) +
geom_hline(yintercept = 0, linetype = "dotted")
data_rescaled <- rescale_mnirs(
data_filtered, ## un-grouped nirs channels to rescale separately
nirs_channels = list(smo2_left, smo2_right),
range = c(0, 100) ## rescale to a 0-100% functional exercise range
)
plot(data_rescaled, time_labels = TRUE) +
geom_hline(yintercept = c(0, 100), linetype = "dotted")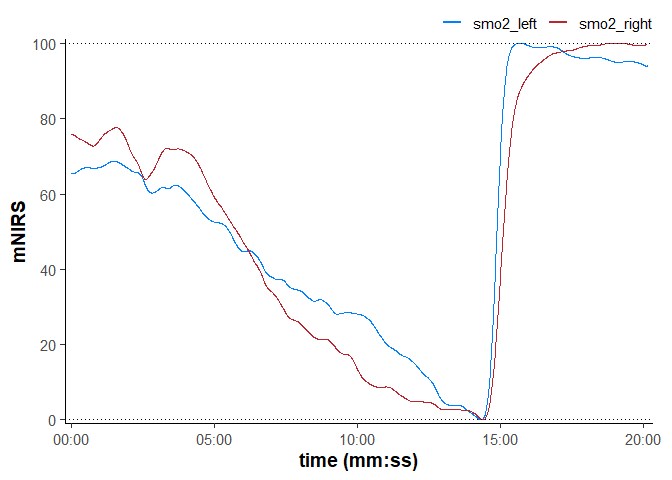
## global option to silence info & warning messages
options(mnirs.verbose = FALSE)
nirs_data <- read_mnirs(
example_mnirs("train.red"),
nirs_channels = c(
smo2_left = "SmO2 unfiltered",
smo2_right = "SmO2 unfiltered"
),
time_channel = c(time = "Timestamp (seconds passed)"),
zero_time = TRUE
) |>
resample_mnirs(method = "linear") |> ## default settings will resample to the same `sample_rate`
replace_mnirs(
invalid_above = 73,
outlier_cutoff = 3,
span = 7
) |>
filter_mnirs(
method = "butterworth",
order = 2,
W = 0.02,
na.rm = TRUE
) |>
shift_mnirs(
nirs_channels = list(smo2_left, smo2_right), ## 👈 channels grouped separately
to = 0,
span = 60,
position = "first"
) |>
rescale_mnirs(
nirs_channels = list(c(smo2_left, smo2_right)), ## 👈 channels grouped together
range = c(0, 100)
)
plot(nirs_data, time_labels = TRUE)
extract_intervals():
detect events and extract intervals## return each interval independently with `event_groups = "distinct"`
distinct <- extract_intervals(
nirs_data, ## channels blank for "distinct" grouping
start = by_time(348, 1064), ## manually identified interval start times
end = by_time(458, 1174), ## interval end time (start + 150 sec)
event_groups = "distinct", ## return a list of data frames for each (2) event
span = c(0, 0), ## no boundary modification
zero_time = FALSE ## return original time values
)
plot(distinct[[1L]], time_labels = TRUE) + plot(distinct[[2L]], time_labels = TRUE)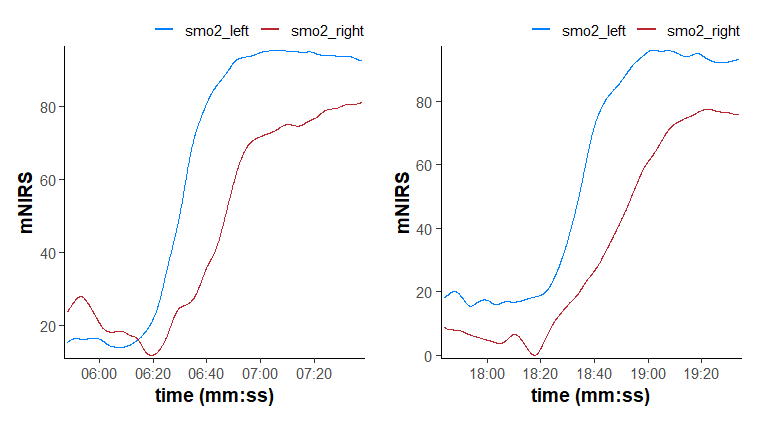
## ensemble average both intervals with `event_groups = "ensemble"`
ensemble <- extract_intervals(
nirs_data, ## channels recycled to all intervals by default
nirs_channels = c(smo2_left, smo2_right),
start = by_time(368, 1084), ## alternatively specify start times + span
event_groups = "ensemble", ## ensemble-average across two intervals
span = c(-20, 90), ## span recycled to all intervals by default
zero_time = TRUE ## re-calculate common time to start from `0`
)
plot(ensemble[[1L]], time_labels = TRUE) +
geom_vline(xintercept = 0, linetype = "dotted")
mnirs
developmentProcess oxygenation kinetics
Monoexponential & sigmoidal non-linear curve fitting
Non-parametric half-response time & slope analysis
Critical oxygenation breakpoint analysis
Oxidative capacity assessment
Repeated occlusion ensemble-averaging and model fitting
Blood volume correction
This package is designed to recognise mNIRS data exported as .csv or .xls(x) files. It should be flexible for use with many different NIRS devices, and compatibility will improve with continued development.
Currently, it has been tested successfully with mNIRS data exported from the following devices and apps:
Generative chatbots are used to assist with code optimisation. All code is thoroughly reviewed and validated by the package author.